Research
The lab investigates the origins of biological diversity, particularly how abrupt genomic events such as polyploidy (genome duplication), chromosomal change, and hybridization have contributed to the evolution and diversity of life. Research integrates computational genomic tools with molecular evolution, phylogenetics, mathematical modeling, field collections, and experimental work across study systems in the Sonoran Desert, Madrean Sky Islands, and global datasets.
1. Macroevolutionary Patterns and Diversification
How does polyploidy influence large-scale patterns of speciation, extinction, and biodiversity accumulation across evolutionary time?
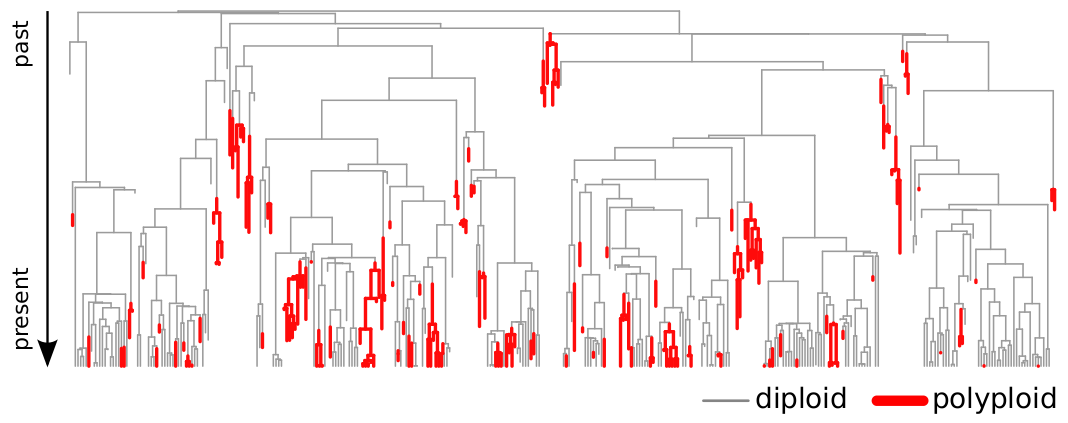
Key Findings:
- Higher speciation AND extinction in polyploid species
- Net diversity increase despite turnover
- Variable patterns across lineages
- Ancient WGDs (whole-genome duplications) shape modern diversity
Methods: Trait-dependent diversification models; Comprehensive ploidy inference; Fossil-calibrated phylogenies; Integrated diversification analyses.
Open Questions: What determines polyploid success versus failure? How do environmental changes affect polyploid diversification? Can we predict rapid diversification potential? How do WGDs interact with other drivers of biodiversity?
Career Opportunities: macroevolutionary modeling, diversification analyses, fossil calibration work, and biodiversity synthesis
2. Phylogenomic Approaches to WGD Evolution
How can we accurately detect and characterize ancient genome duplications across the tree of life? What do large-scale phylogenomic analyses reveal about the timing, frequency, and evolutionary consequences of WGDs?
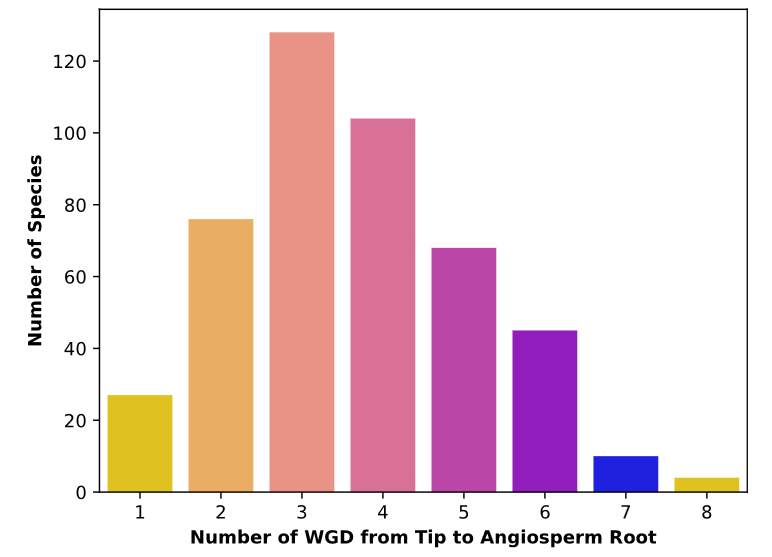
Key Findings:
- Complex duplication histories across land plants
- Phylogenetic uncertainty affects WGD inference
- Non-random WGD distribution across clades
- Multiple genomic features improve detection accuracy
Methods: Large-scale phylogenomic analysis; Uncertainty-aware WGD inference; Synteny and gene tree integration; Machine learning detection algorithms (SLEDGE).
Key References:
- One Thousand Plant Transcriptomes Initiative (2019, Nature)
- McKibben et al. 2024, American Journal of Botany
- Li et al. 2015, Science Advances
- Barker et al. 2008, MBE
- Barker et al. 2016, AJB
- Mabry et al. 2020, AJB
Career Opportunities: large-scale data analysis, phylogenetic methods, computational biology, and collaborative projects with international consortia
3. Diploidization and Post-Polyploid Genome Evolution
Why do polyploid species diploidize if they were successful as polyploids? Does natural selection or drift drive diploidization? Are there universal rules that govern genome evolution following genome duplication?
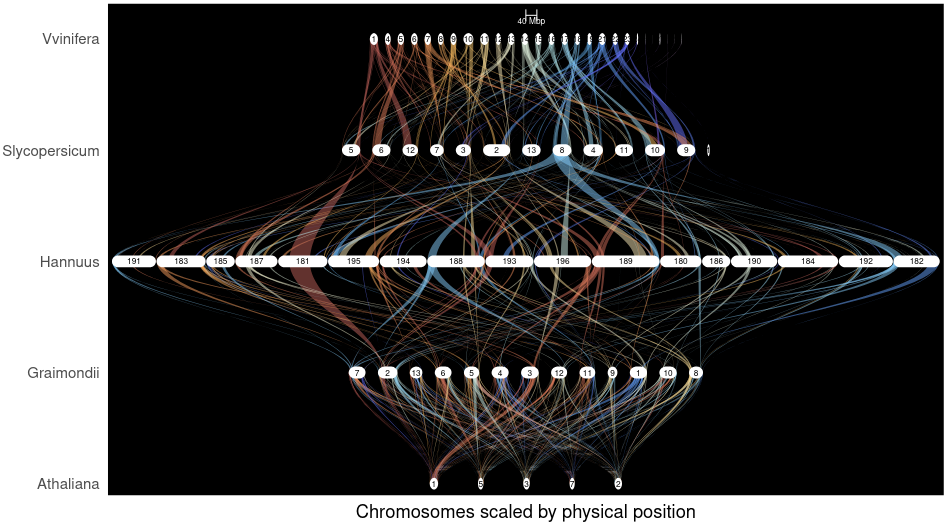

Key Findings:
- Variable diploidization rates across lineages create genomic diversity
- Different processes of diploidization and post-polyploid genome evolution proceed independently
- Gene retention shows both stochastic and selective patterns
- Synteny conservation varies dramatically between plant groups
Methods: Comparative fractionation analysis; Machine learning classification of WGD paralogs (Frackify); Synteny-based analyses (SynTRACE); Phylogenomic integration approaches.
Notable Research:
Career Opportunities: computational phylogenomics, comparative genomics, and collaborative projects with plant genome consortia
4. Chromosome Number Evolution
Why do rates of chromosome gain and loss vary dramatically across the phylogeny? What evolutionary forces drive the gain and loss of chromosomes, and ultimately the evolution of chromosome number?
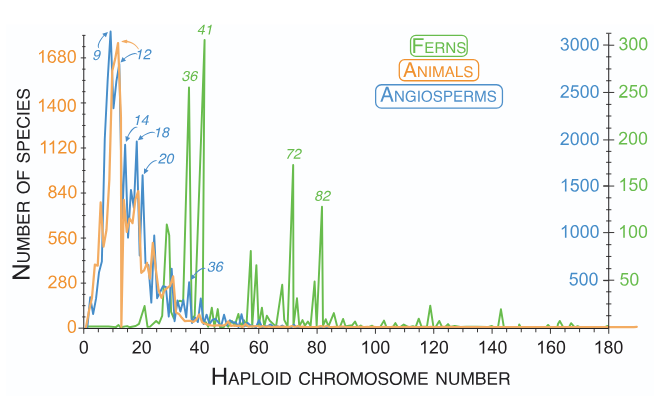
Key Findings:
- Meiotic drive profoundly impacts chromosome evolution
- High numbers of chromosomes in ferns and rapid loss in angiosperms may both be explained by drive
- Annual plants show 11.41x higher chromosome loss rates
- No correlation between chromosome number and reproductive barriers
Methods: Comparative karyotype analysis; Analyses to test for signatures of meiotic drive; Extreme systems (Xanthisma n = 2); ChromEvol and BiChroM modeling.
Notable Research:
- Román-Palacios et al. 2021, Journal of Evolutionary Biology
- Plačková et al. 2024
- Kinosian & Barker 2025, Molecular Ecology
Career Opportunities: field collections, cytological work, theoretical modeling, and comparative analyses
5. Population Genomics of Polyploid Systems
How do multiple origins of polyploid species contribute to polyploid success? How do the population genetics of polyploid species influence their macroevolutionary outcomes?
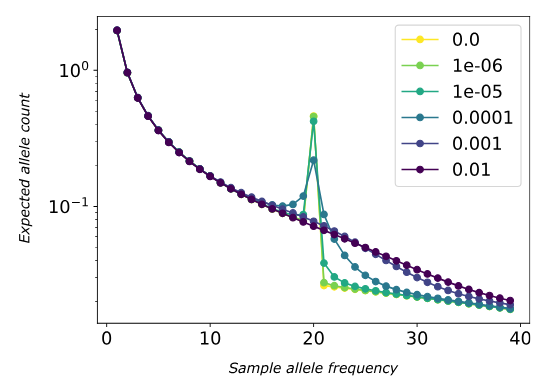
Key Findings:
- Complex inheritance creates unique evolutionary trajectories
- Multiple origins common in polyploid formation
- Homoeolog exchange affects selection efficacy
- Demographic patterns differ from diploid expectations
Methods: Polyploid demographic modeling; Low-coverage SNP calling methods; Inheritance pattern estimation; Complex mating system analysis.
Notable Research:
Career Opportunities: population genetics theory, statistical method development, field sampling, and genomic data analysis
6. Climate Adaptation and Ecological Genomics
How does polyploidy influence rates of adaptation and speciation? Do polyploid species demonstrate greater ecological plasticity than their diploid relatives?
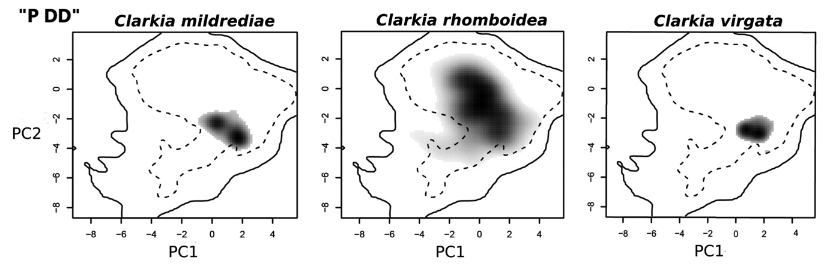
Key Findings:
- Accelerated multivariate niche evolution in polyploids
- Genome duplication facilitates novel niche occupation
- Enhanced stress response potential
- Co-evolutionary advantages through gene duplication
Methods: Ecological niche modeling; Environmental gradient sampling; Comparative phylogenetic approaches; Genomic-ecological data integration.
Notable Research:
Career Opportunities: field work in the Sonoran Desert and Sky Islands, ecological modeling, conservation genomics, and climate change research
7. Agricultural Genomics and Crop Domestication
How do ancient genome duplications continue to influence contemporary agriculture? Can evolutionary genomics inform crop improvement strategies?
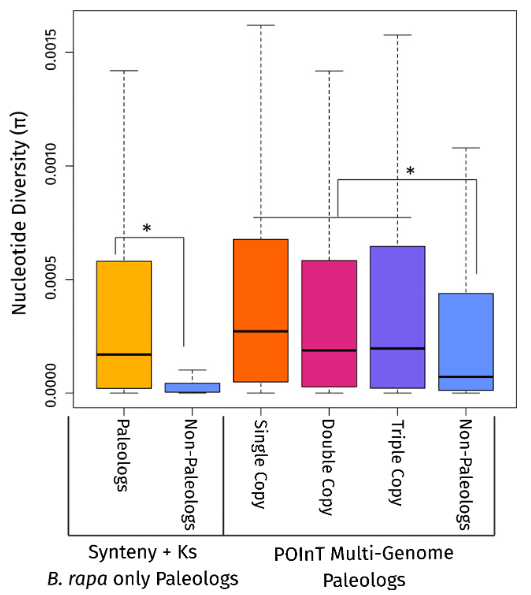
Key Findings:
- Paleolog enrichment in domestication genes across many crops
- Higher genetic diversity in paralogs retained from ancient WGDs
- Single-copy paleologs most enriched
- Ancient WGDs enable modern crop improvement
Methods: Machine learning gene classification (Frackify); Population genomics of crop diversity; Phylogenetic comparative approaches; Selection analyses.
Notable Research:
Career Opportunities: computational genomics, agricultural collaborations, molecular evolution analyses, and applied research with societal impact
8. Bioinformatic Tool Development
How can machine learning and computational approaches detect complex genomic patterns that traditional methods cannot capture?
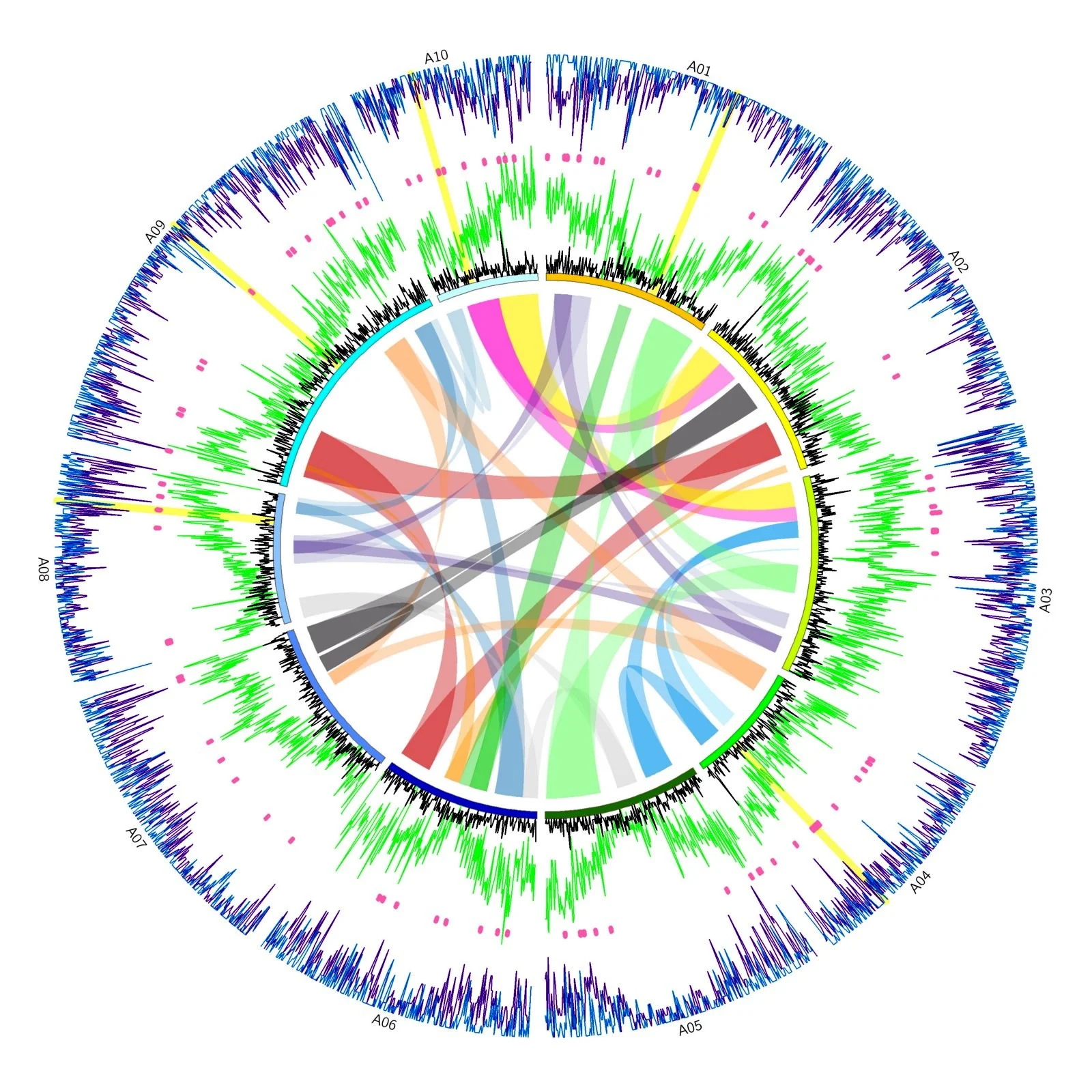
Key Tools:
- Frackify — gradient boosted trees for gene classification
- HyDe-CNN — neural networks for hybrid detection
- SynTRACE — synteny-based genome reconstruction
- Ploidify — probabilistic ploidy inference
- GOgetter & GOmosaic — polyploid functional annotation
Career Opportunities: machine learning, software development, open-source contributions, and computational methods training
Joining Our Research Program
If you are interested in exploring these evolutionary questions about genome duplications, polyploidy, and biodiversity, we welcome you to join our lab. Please visit our Contact page for more information about current opportunities and how to apply.