Code
Most code below is available at https://gitlab.com/barker-lab unless otherwise indicated below.
EvoPipes
A repository of the core EvoPipes pipelines is available at https://gitlab.com/barker-lab/EvoPipes. This is a suite of bioinformatic tools for whole genome duplication inference, HMM-based gene translation, and ortholog identification.

Frackify
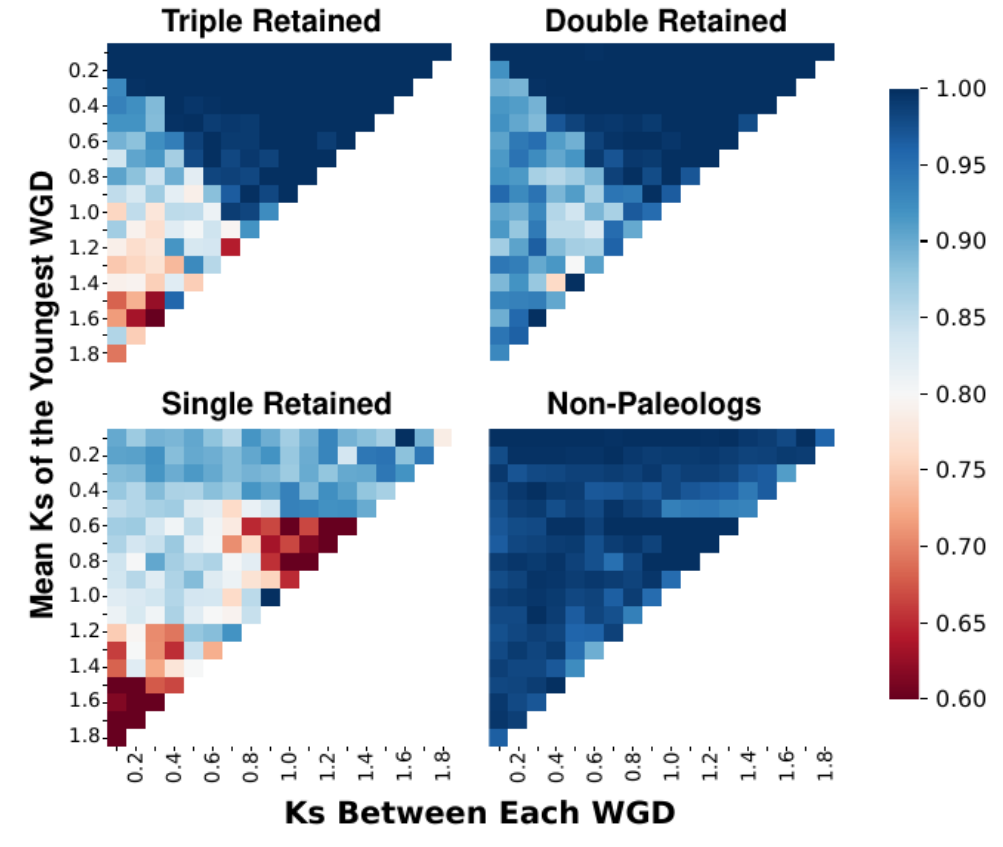
Machine learning classification of gene duplication origins using gradient boosted decision trees and XGBoost. Combines synteny and divergence features to identify paleologs…
Ploidify
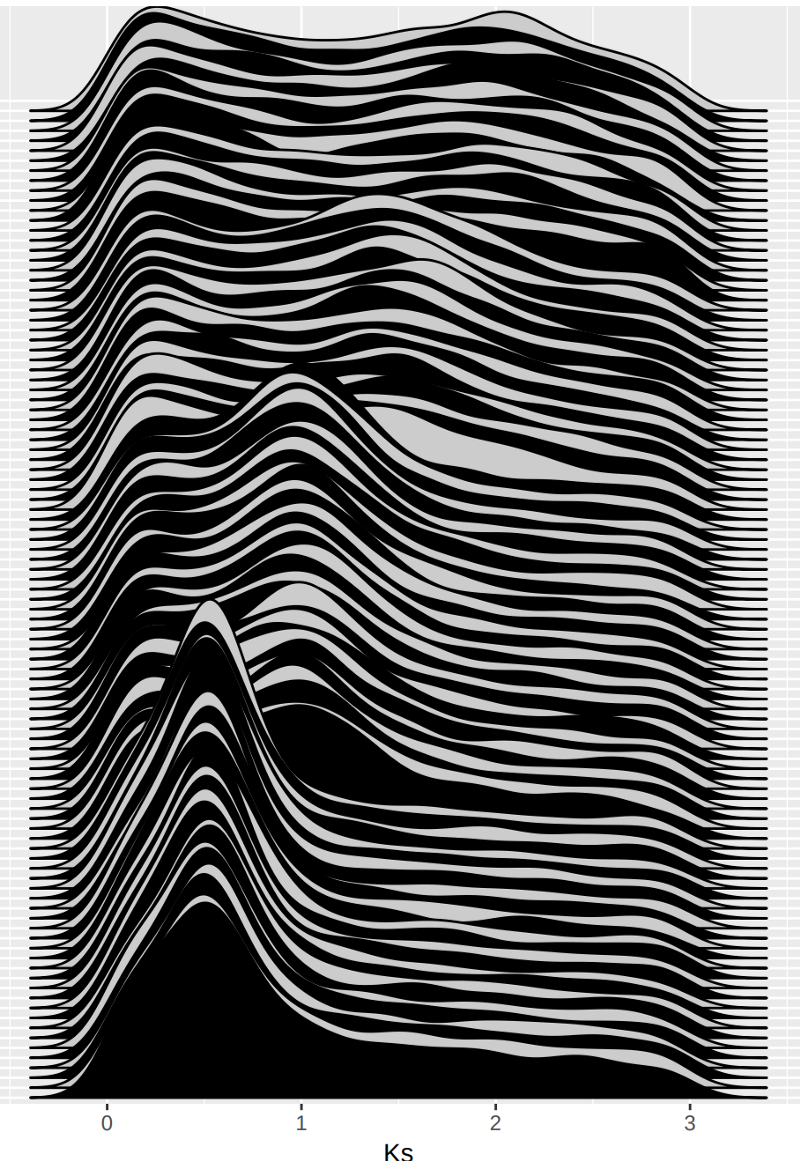
Logistic regression model for identifying ploidal levels of ancient whole genome duplications using Frackify output data.

SLEDGE
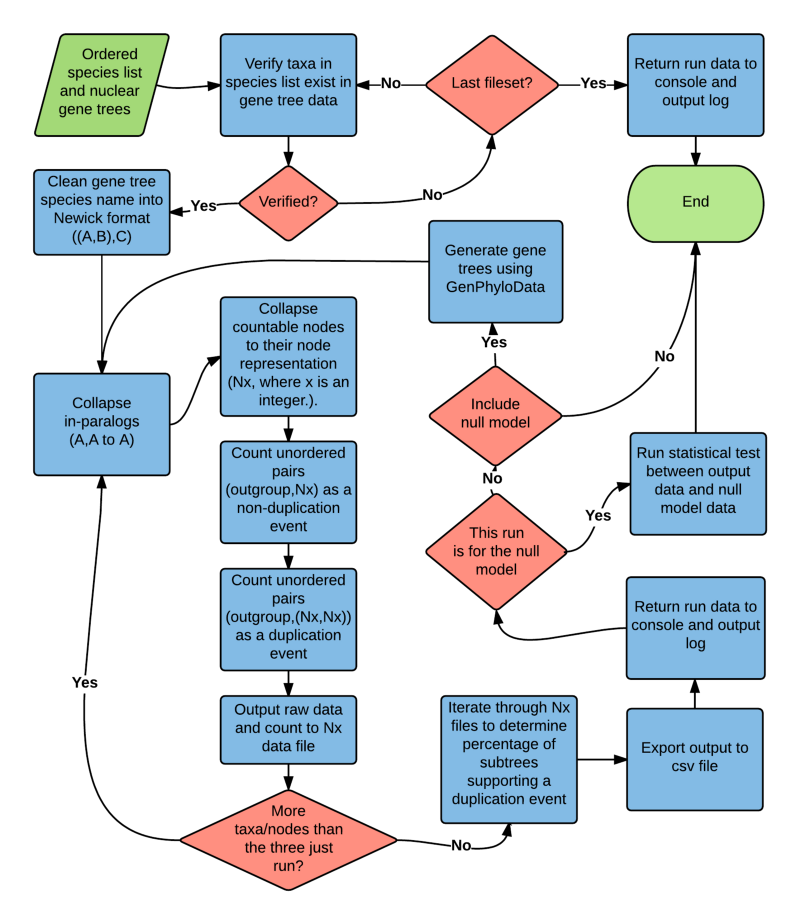
Machine learning tool for automated detection and classification of ancient whole genome duplications in paralog Ks distributions.
HyDe-CNN
Convolutional neural network approaches for classifying hybridization events and hybrid speciation patterns in genomic data.
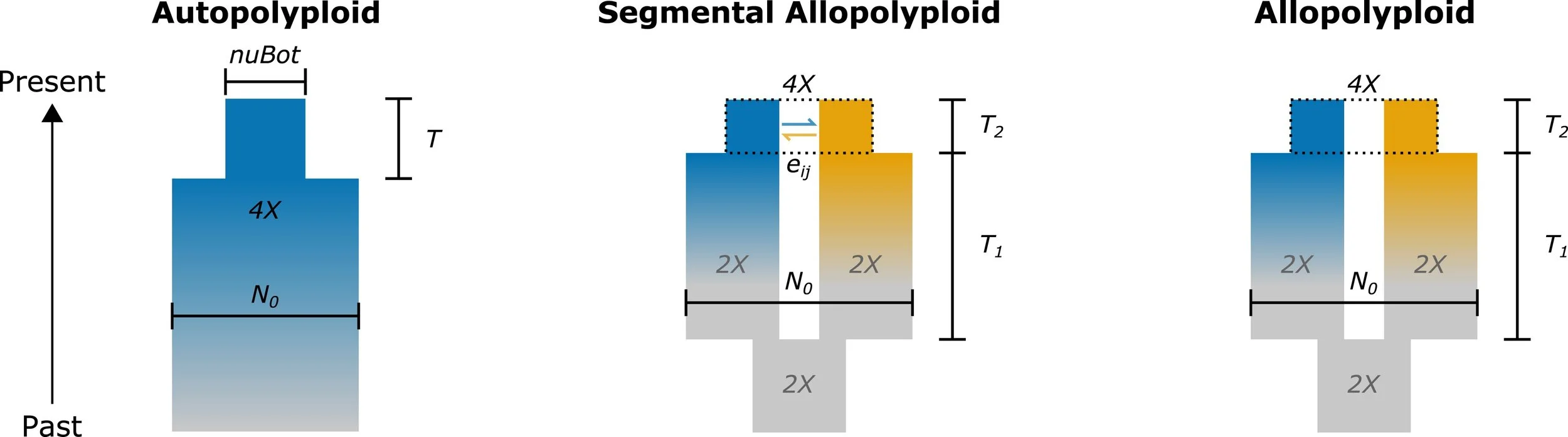
dadi Models for Polyploids
Extended demographic inference models for polyploid and inbred organisms, developed by Paul Blischak.
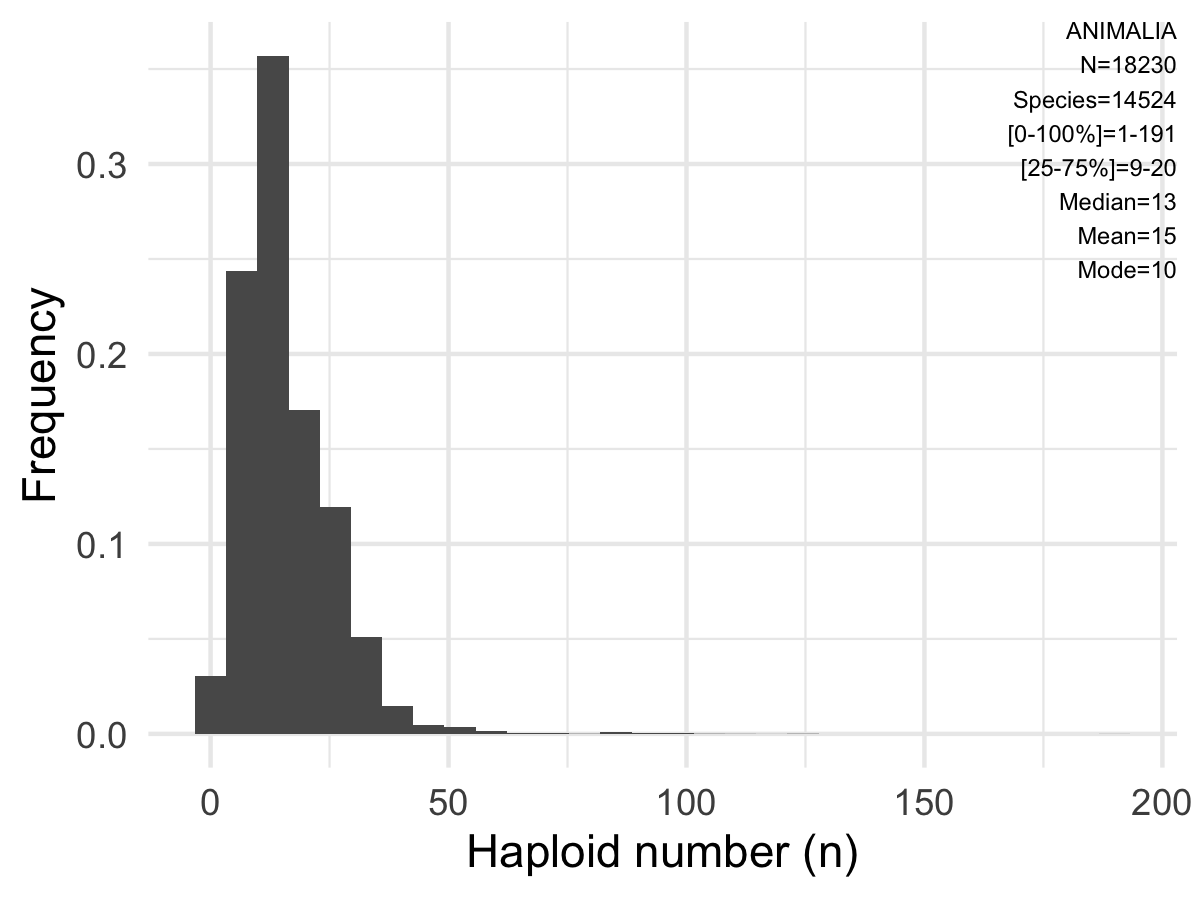
Animal Chromosome Count Database (ACC)
The largest database of animal chromosomal counts…curated chromosome numbers across the animal Tree of Life.
- GitHub: https://github.com/cromanpa94/ACC
- Web interface: https://cromanpa94.github.io/ACC/
GOgetter
Functional annotation tool for analyzing gene ontology patterns in plant genomes.
In Development
- SynTRACE — GPU-accelerated syntenic network analysis
- GOmosaic — Enhanced GO term visualization tool
Legacy Tools
- MAPS
- NU-IN
- SCARF